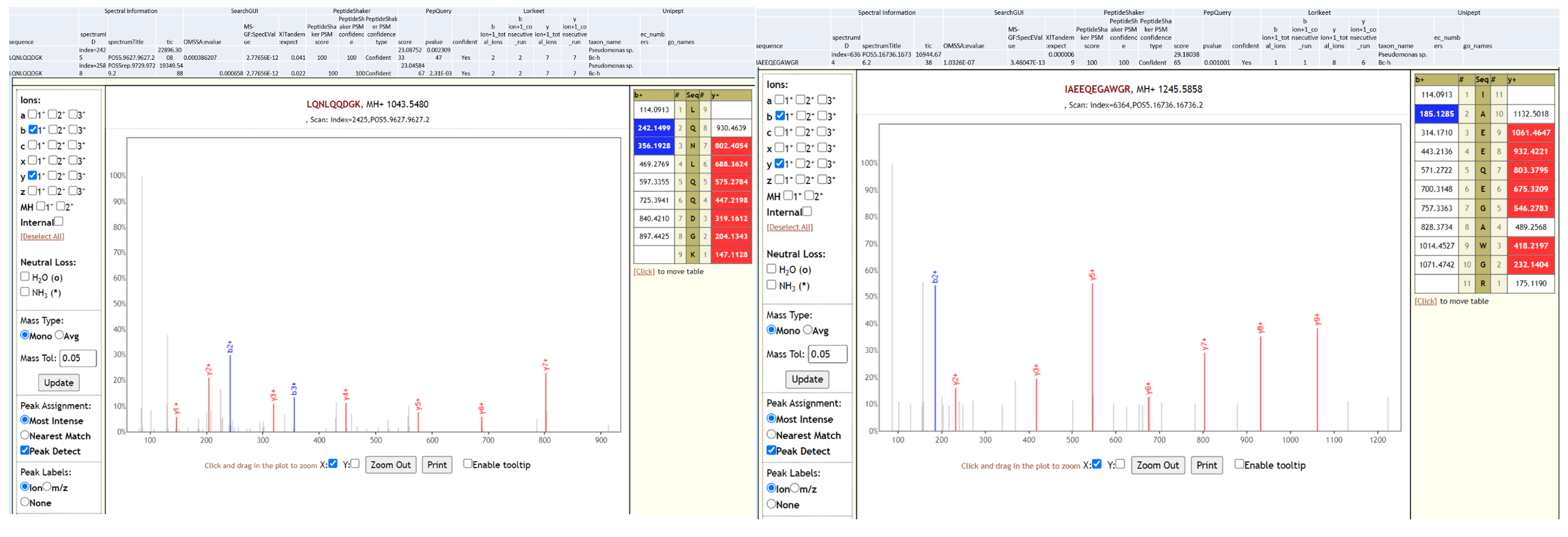
Given that the spectrum number/index is not always reliable, and given that there is no one standard format for how to annotate spectrum number/index across all the different spectrum file formats, PeptideShaker basically ignores the spectrum numbers, and instead relies on the spectrum titles. 
What needs to be in the MGF header to make this mapping work properly? I notice in the 'default_PSM_report' from Peptide Shaker that it doesn't seem able to get the actual spectrum number out for the 'Spectrum Number' column. Well, unless you pass the first of them through Comet that is. (Also should not include non-standard characters etc., but I guess that is a given.) Both of the headers above should therefore be fine for both SearchGUI and PeptideShaker. Our main requirement is that the spectrum title is unique and can thus be used to uniquely identify a spectrum. On a side note, what would an ideal MGF header include for SearchGUI and PeptideShaker look like? And in what format? For example, does anything about the first MGF header I posted above make you think it might give an issue with this mapping you mention? I am using SearchCLI with the below input command to generate these data: Thu Sep 20 09:07: PeptideShaker Processing Canceled. : Index: 15129, Size: 15129Īt (ArrayList.java:657)Īt (ArrayList.java:433)Īt .io.(MgfIndex.java:239)Īt .(SpectrumFactory.java:1114)Īt eu.$IdProcessorFromFile.importSpectrum(FileImporter.java:999)Īt eu.$IdProcessorFromFile.importPsms(FileImporter.java:720)Īt eu.$IdProcessorFromFile.importFiles(FileImporter.java:476)Īt eu.importFiles(FileImporter.java:151)Īt eu.(PeptideShaker.java:229)Īt eu.createProject(PeptideShakerCLI.java:738)Īt eu.call(PeptideShakerCLI.java:218)Īt eu.main(PeptideShakerCLI.java:938) Thu Sep 20 09:07: An error occurred while loading the identification files: Thu Sep 20 09:07: Loading spectra for ch_.xml. Thu Sep 20 09:07: Reading identification files. Thu Sep 20 09:07: Establishing local database connection. Thu Sep 20 09:07: Importing gene mappings. Thu Sep 20 09:07: FASTA file import completed. Thu Sep 20 09:06: Importing sequences from uniprot_human-crap_sep2018_FWD_concatenated_target_decoy.fasta. Thu Sep 20 09:06: Import process for fractions (Sample: ch_XXXXX.raw_searchgui_out.zip, Replicate: 1) Thu Sep 20 09:06: Unzipping ch_XXXXX.raw_searchgui_out.zip.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
